What Determines Primary Structure Of A Protein
Muz Play
Apr 03, 2025 · 6 min read
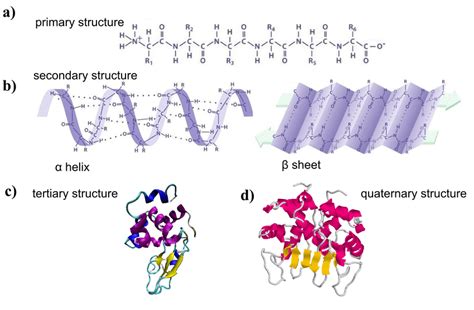
Table of Contents
What Determines the Primary Structure of a Protein?
The primary structure of a protein, the fundamental building block upon which all higher-level structures are built, is nothing short of remarkable. It's the unique linear sequence of amino acids linked together by peptide bonds. This seemingly simple arrangement dictates the protein's ultimate three-dimensional shape and, consequently, its function. Understanding what determines this primary structure is crucial to grasping the intricacies of protein biology and its role in various biological processes.
The Amino Acid Alphabet: The Building Blocks of Proteins
Before diving into the determinants of primary structure, let's briefly revisit the fundamental components: amino acids. Proteins are polymers composed of monomers called amino acids. There are 20 standard amino acids, each possessing a unique side chain (R-group) attached to a central carbon atom. This R-group imparts specific chemical properties to each amino acid, influencing how it interacts with other amino acids and the surrounding environment. These properties include:
- Hydrophobicity/Hydrophilicity: Some amino acids have hydrophobic (water-repelling) side chains, while others are hydrophilic (water-attracting). This property plays a crucial role in protein folding and stability.
- Charge: Some amino acids possess charged side chains (positive or negative), contributing to electrostatic interactions within the protein and with its surroundings.
- Size and Shape: The size and shape of the R-group influence steric hindrance and packing within the protein's three-dimensional structure.
- Reactivity: Certain amino acids have reactive side chains that can participate in covalent modifications or catalytic reactions.
The Peptide Bond: Linking Amino Acids Together
The amino acids are linked together through a peptide bond, a covalent bond formed between the carboxyl group (-COOH) of one amino acid and the amino group (-NH2) of another. This reaction releases a water molecule, a process known as dehydration synthesis. The resulting chain of amino acids is called a polypeptide chain. The peptide bond itself possesses a partial double bond character due to resonance, restricting rotation around the bond and influencing the protein's conformation.
Genetic Code: The Blueprint for Protein Synthesis
The primary structure of a protein is ultimately dictated by the genetic code. This code is embedded within the DNA sequence of a gene. Each gene carries the instructions for synthesizing a specific polypeptide chain. The DNA sequence is transcribed into messenger RNA (mRNA), which then serves as a template for protein synthesis during translation.
Transcription: DNA to mRNA
The process of transcription involves the synthesis of an mRNA molecule that is complementary to the DNA sequence of a gene. This mRNA molecule carries the genetic information from the nucleus to the ribosomes in the cytoplasm, where protein synthesis occurs.
Translation: mRNA to Protein
Translation is the process where the mRNA sequence is decoded by ribosomes to synthesize the polypeptide chain. Ribosomes read the mRNA sequence in codons (three-nucleotide sequences). Each codon specifies a particular amino acid. Transfer RNA (tRNA) molecules, each carrying a specific amino acid, recognize and bind to their corresponding codons on the mRNA. The ribosome facilitates the formation of peptide bonds between the amino acids, creating the growing polypeptide chain. The process continues until a stop codon is encountered, signaling the termination of translation.
Factors Influencing Primary Structure Beyond the Genetic Code
While the genetic code provides the primary blueprint, other factors can subtly influence the final primary structure:
Post-Translational Modifications
After translation, the newly synthesized polypeptide chain can undergo various post-translational modifications that alter its primary structure. These modifications include:
- Glycosylation: The addition of carbohydrate groups.
- Phosphorylation: The addition of phosphate groups.
- Acetylation: The addition of acetyl groups.
- Proteolytic Cleavage: The removal of amino acid segments.
These modifications can significantly affect the protein's function, stability, and interactions with other molecules. For example, glycosylation can affect protein solubility and cell-surface recognition, while phosphorylation can regulate protein activity.
Genetic Mutations: Variations in the Code
Mutations in the DNA sequence can lead to changes in the amino acid sequence of the resulting protein. These changes can be:
- Missense mutations: A single nucleotide change leading to a different amino acid being incorporated. This can have varying effects depending on the location and nature of the amino acid change. Some missense mutations are silent (causing no significant change in protein function), while others can significantly alter protein function or stability.
- Nonsense mutations: A nucleotide change leading to a premature stop codon, resulting in a truncated protein. These mutations often lead to non-functional proteins.
- Frameshift mutations: Insertions or deletions of nucleotides that shift the reading frame of the mRNA, leading to a completely altered amino acid sequence downstream from the mutation.
Environmental Factors: External Influences
Environmental factors can also subtly influence protein synthesis and potentially lead to variations in the primary structure, although this is less direct than genetic factors. For example, stress conditions (heat shock, oxidative stress) can induce changes in gene expression, leading to variations in the amount or even specific isoforms of proteins produced. Similarly, nutrient availability can affect post-translational modifications.
The Importance of Primary Structure in Determining Higher-Level Structures
The primary structure doesn't just exist in isolation; it's the foundation upon which the secondary, tertiary, and quaternary structures are built. The amino acid sequence dictates how the polypeptide chain folds into its functional three-dimensional shape.
Secondary Structure: Local Folding Patterns
The primary structure influences the formation of secondary structures, local folding patterns stabilized by hydrogen bonds between the backbone atoms of the polypeptide chain. These patterns include:
- Alpha-helices: Right-handed coiled structures.
- Beta-sheets: Extended polypeptide chains arranged in parallel or antiparallel configurations.
- Turns and Loops: Connecting elements between alpha-helices and beta-sheets.
The distribution and arrangement of amino acids along the polypeptide chain dictate which secondary structures are energetically favorable. For instance, proline residues often disrupt alpha-helices, while glycine residues are flexible and found in turns.
Tertiary Structure: Overall 3D Shape
The tertiary structure represents the overall three-dimensional arrangement of the polypeptide chain, stabilized by a variety of non-covalent interactions including:
- Hydrophobic interactions: Clustering of hydrophobic amino acids in the protein's core.
- Hydrogen bonds: Interactions between polar side chains and the polypeptide backbone.
- Ionic bonds (salt bridges): Electrostatic interactions between charged amino acid side chains.
- Disulfide bonds: Covalent bonds formed between cysteine residues.
The spatial arrangement of secondary structure elements within the tertiary structure is primarily dictated by the primary sequence and interactions between amino acid side chains. This structure is often crucial for the protein's function; for instance, the active site of an enzyme is typically formed within a specific tertiary structure arrangement.
Quaternary Structure: Multiple Polypeptide Chains
Some proteins consist of multiple polypeptide chains (subunits) assembled together to form a functional protein complex. This arrangement constitutes the quaternary structure. The interaction between subunits is determined by the primary structure of each subunit, as specific interaction surfaces are created by the folding of individual chains. The primary structures of the subunits thus dictate the overall architecture and function of the quaternary structure.
Conclusion: A Delicate Balance
The primary structure of a protein, while seemingly simple – a linear sequence of amino acids – is a complex and precisely determined sequence that forms the basis of all higher-level structural features and functions. The genetic code dictates the initial sequence, but post-translational modifications and environmental factors can further influence the final product. Understanding these determinants is pivotal to unraveling the intricacies of protein biology, developing new therapies targeting misfolded proteins associated with disease, and designing new proteins with customized properties for biotechnology applications. The remarkable precision of this process highlights the exquisite control mechanisms operating within cells to ensure the faithful synthesis and functional assembly of these crucial biological molecules.
Latest Posts
Latest Posts
-
Example Of Stoichiometry In Real Life
Apr 04, 2025
-
Dna Biology And Technology Dna And Rna Structure
Apr 04, 2025
-
Narcotics Act On The Central Nervous System By Producing A
Apr 04, 2025
-
Two Plants Heteroygous For Pod Color Are Crossed Refer To
Apr 04, 2025
-
What Two Main Products Result From Photosynthesis
Apr 04, 2025
Related Post
Thank you for visiting our website which covers about What Determines Primary Structure Of A Protein . We hope the information provided has been useful to you. Feel free to contact us if you have any questions or need further assistance. See you next time and don't miss to bookmark.
