What Is Sigma Factor In Transcription
Muz Play
Apr 05, 2025 · 7 min read
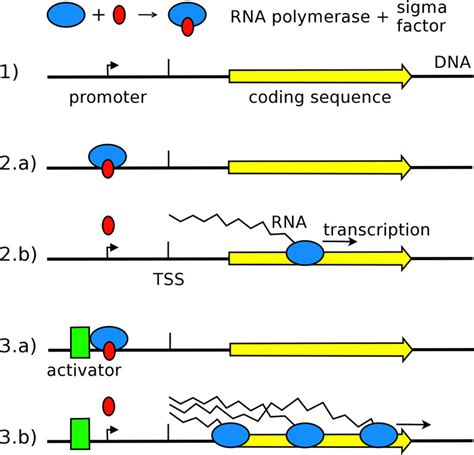
Table of Contents
What is Sigma Factor in Transcription? A Deep Dive into Bacterial RNA Polymerase
Transcription, the process of synthesizing RNA from a DNA template, is fundamental to life. In bacteria, this crucial process is orchestrated by RNA polymerase, a complex enzyme whose activity is intricately regulated by a protein subunit called the sigma factor (σ factor). Understanding sigma factors is key to comprehending bacterial gene expression and its implications for various biological processes, including virulence, stress response, and adaptation.
The Role of Sigma Factors in Bacterial Transcription Initiation
Bacterial RNA polymerase is a holoenzyme comprised of a core enzyme and a sigma factor. The core enzyme consists of five subunits: α2ββ'ω. This core enzyme possesses the catalytic activity necessary for RNA synthesis, but it lacks the ability to specifically recognize and bind to promoter regions on the DNA, the specific sequences that signal the start of a gene. This is where the sigma factor comes into play.
The sigma factor acts as a crucial recognition and binding subunit. It directs the core enzyme to specific promoter regions on the DNA, enabling the initiation of transcription at the correct location. Without a sigma factor, the core enzyme binds non-specifically to DNA, resulting in inefficient and indiscriminate transcription. The sigma factor's interaction with the core enzyme temporarily transforms it into a fully functional holoenzyme capable of initiating transcription. Think of the sigma factor as the GPS for the RNA polymerase, guiding it to the precise location where it needs to begin transcribing.
How Sigma Factors Recognize Promoters: The Consensus Sequence
Promoters are typically located upstream of the transcription start site (+1 site) and consist of conserved sequences, known as consensus sequences, which are recognized by the sigma factor. The most common consensus sequences are the -10 and -35 regions, located approximately 10 and 35 base pairs upstream of the +1 site, respectively. These sequences are not identical across all genes; variations in the promoter sequence affect the strength of promoter binding and, consequently, the frequency of transcription initiation.
The -10 region, also known as the Pribnow box, has a consensus sequence of TATAAT. The -35 region typically has a consensus sequence of TTGACA. The sigma factor interacts with these sequences through specific amino acid residues within its protein structure. The strength of the interaction is influenced by the degree of similarity between the promoter sequence and the consensus sequence. Strong promoters have sequences that closely match the consensus sequences, leading to higher transcription rates, while weak promoters exhibit greater divergence, resulting in lower transcription rates.
The Transcription Initiation Cycle: A Step-by-Step Process
The process of transcription initiation involving sigma factors can be broken down into several key steps:
-
Holoenzyme Formation: The sigma factor binds to the core RNA polymerase, forming the holoenzyme.
-
Promoter Recognition and Binding: The sigma factor within the holoenzyme recognizes and binds to the -35 and -10 regions of the promoter sequence on the DNA. This binding involves several protein-DNA interactions, establishing a closed promoter complex.
-
Promoter Melting: The holoenzyme unwinds a short stretch of DNA around the -10 region, forming an open promoter complex. This unwinding process requires energy and is facilitated by the sigma factor.
-
Initiation of RNA Synthesis: RNA polymerase initiates the synthesis of RNA by adding the first nucleotide to the 5' end of the growing RNA molecule.
-
Promoter Escape: Once a short RNA transcript (around 8-10 nucleotides) is synthesized, the sigma factor dissociates from the core enzyme. This step signifies the transition from initiation to elongation. The core enzyme then continues transcribing the gene, elongating the RNA molecule.
The Diversity of Sigma Factors: Adapting to Different Environmental Conditions
Bacteria do not rely on a single sigma factor; they often possess a family of sigma factors, each recognizing a specific set of promoters and regulating the expression of a particular set of genes. This diversity allows bacteria to respond effectively to various environmental conditions and developmental cues.
The primary sigma factor, denoted as σ70 (the number refers to its molecular weight in kilodaltons), is responsible for transcribing the majority of "housekeeping" genes, those necessary for basic cellular functions under normal growth conditions. However, under stress or during specific developmental stages, alternative sigma factors are activated.
Examples of Alternative Sigma Factors and their Roles:
-
σ32 (heat shock sigma factor): This sigma factor is induced by heat shock and other forms of cellular stress, initiating the transcription of heat shock proteins that protect the cell from damage.
-
σ54 (nitrogen regulation sigma factor): This sigma factor regulates genes involved in nitrogen metabolism, responding to nitrogen limitation in the environment.
-
σS (general stress response sigma factor): This sigma factor is activated under various stress conditions, including starvation, osmotic shock, and oxidative stress, regulating the expression of genes involved in stress response and survival.
-
σB (sporulation sigma factor): This sigma factor is crucial for the sporulation process in Bacillus subtilis, initiating the transcription of genes required for the formation of spores.
-
Extracytoplasmic function (ECF) sigma factors: This large family of sigma factors responds to a wide range of environmental stimuli and stress conditions, often related to the cell envelope. Their regulatory roles cover a diverse array of functions, including response to antibiotics, changes in osmolarity, and alterations in cell wall integrity.
The expression of alternative sigma factors is highly regulated, ensuring that genes are transcribed only when appropriate. This regulation often involves complex signaling pathways that sense environmental changes and activate or repress the synthesis or activity of specific sigma factors.
Regulation of Sigma Factor Activity: A Multifaceted Control Mechanism
The activity of sigma factors is tightly controlled at multiple levels to ensure precise regulation of gene expression. These mechanisms include:
-
Transcriptional regulation: The expression of sigma factor genes themselves can be controlled by other regulatory proteins, ensuring the appropriate levels of sigma factors are present under specific conditions.
-
Post-translational modifications: Sigma factors can undergo post-translational modifications, such as phosphorylation or proteolysis, which affect their activity and stability.
-
Anti-sigma factors: Some sigma factors are regulated by anti-sigma factors, which bind to the sigma factor and inhibit its interaction with the core RNA polymerase. The release of the sigma factor from the anti-sigma factor is often triggered by environmental stimuli or signaling pathways.
-
Sigma factor chaperones: Some sigma factors are maintained in an inactive state through interactions with chaperone proteins. The release of the sigma factor from the chaperone is required for its activation.
These regulatory mechanisms, acting in concert, ensure a fine-tuned control of gene expression, allowing bacteria to adapt to diverse environments and maintain homeostasis.
Sigma Factors in Pathogenesis and Antibiotic Resistance
The role of sigma factors extends beyond fundamental cellular processes. They play a crucial role in bacterial pathogenesis and the development of antibiotic resistance. Many bacterial pathogens utilize alternative sigma factors to regulate the expression of virulence factors, proteins that contribute to the pathogen's ability to cause disease.
For example, the sigma factor σE in Pseudomonas aeruginosa regulates genes involved in the production of biofilm, an important virulence factor. Similarly, the sigma factor σB in Staphylococcus aureus regulates genes involved in stress resistance and virulence factor expression. Understanding the roles of these sigma factors in pathogenesis is critical for developing strategies to combat bacterial infections.
Furthermore, some sigma factors are implicated in the development of antibiotic resistance. The expression of genes encoding efflux pumps, which remove antibiotics from the bacterial cell, can be regulated by sigma factors. Understanding how sigma factors contribute to antibiotic resistance is crucial for developing novel antimicrobial therapies.
Conclusion: Sigma Factors – Master Regulators of Bacterial Gene Expression
Sigma factors are essential components of the bacterial transcription machinery, serving as master regulators of gene expression. Their ability to recognize and bind to specific promoter sequences, coupled with their diverse and tightly regulated nature, enables bacteria to respond effectively to a wide range of environmental cues and developmental signals. Their involvement in fundamental cellular processes, as well as pathogenesis and antibiotic resistance, makes them attractive targets for the development of novel antimicrobial strategies. Further research into the intricate regulatory mechanisms governing sigma factor activity will undoubtedly continue to unravel their vital role in bacterial biology and pave the way for new therapeutic interventions.
Latest Posts
Latest Posts
-
Is Orange Juice An Acid Or Base
Apr 05, 2025
-
Which Is Smaller An Atom Or A Molecule
Apr 05, 2025
-
Stopping A Filibuster Requires That
Apr 05, 2025
-
What Are The Two Major Divisions Of The Skeletal System
Apr 05, 2025
-
Delta S Delta H Delta G Chart
Apr 05, 2025
Related Post
Thank you for visiting our website which covers about What Is Sigma Factor In Transcription . We hope the information provided has been useful to you. Feel free to contact us if you have any questions or need further assistance. See you next time and don't miss to bookmark.
