What Is The Base Pairing Rule For Rna
Muz Play
Apr 05, 2025 · 5 min read
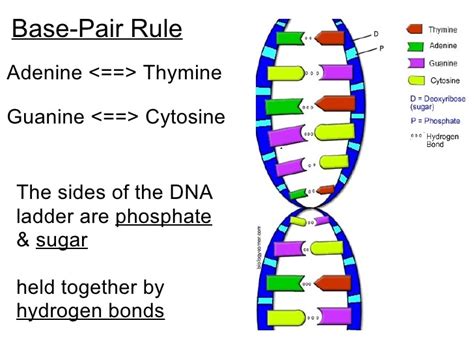
Table of Contents
What is the Base Pairing Rule for RNA?
RNA, or ribonucleic acid, is a crucial molecule in all living cells, playing a vital role in protein synthesis and various other cellular processes. Understanding its structure, particularly the base pairing rules, is fundamental to grasping its function and significance in molecular biology. While DNA famously uses the base pairing rule of adenine (A) with thymine (T) and guanine (G) with cytosine (C), RNA exhibits a slightly different pattern due to its single-stranded nature and the presence of uracil (U) instead of thymine. This article delves deep into the intricacies of RNA base pairing, exploring its variations, exceptions, and implications in different RNA structures and functionalities.
The Fundamental Base Pairing Rule in RNA
The fundamental base pairing rule for RNA is:
- Adenine (A) pairs with Uracil (U)
- Guanine (G) pairs with Cytosine (C)
This rule governs the formation of secondary structures within RNA molecules. Unlike DNA's double helix, RNA is typically single-stranded. However, this single strand often folds back on itself, forming internal base pairs, leading to complex three-dimensional structures crucial for their biological roles. The A-U and G-C base pairs are stabilized by hydrogen bonds: two hydrogen bonds between A and U, and three hydrogen bonds between G and C. This difference in hydrogen bond number contributes to the differing stability of these base pairs.
Variations and Exceptions to the Base Pairing Rule
While the A-U and G-C pairings are the primary rules, several exceptions and variations exist, particularly in non-canonical base pairing interactions. These exceptions often contribute to the structural diversity and functionality of RNA molecules.
Wobble Base Pairing
Wobble base pairing describes non-Watson-Crick base pairing, where less stringent hydrogen bonding occurs between bases. This is particularly relevant in the context of tRNA (transfer RNA) molecules, where the anticodon loop interacts with the mRNA (messenger RNA) codon during translation. Wobble base pairing allows a single tRNA to recognize multiple codons, expanding the decoding capacity of the translational machinery. Examples include:
- G-U wobble pairs: Guanine can pair with uracil through a single hydrogen bond.
- I-U, I-A, I-C wobble pairs: Inosine (I), a modified base found in tRNA, can pair with U, A, and C. This flexibility is crucial for the degeneracy of the genetic code.
Non-Watson-Crick Base Pairs
Beyond wobble pairing, other non-Watson-Crick base pairs can form within RNA structures. These interactions are often crucial for stabilizing complex tertiary structures, loops, and bulges. Examples include:
- A-A base pairs: These can form under specific conditions, contributing to RNA structural stability.
- G-G base pairs: Similar to A-A pairs, G-G base pairs can contribute to structural stability, particularly in specific RNA motifs.
- U-U base pairs: Less common than other non-canonical pairs, U-U pairs can still be observed in certain RNA structures.
Importance of Base Pairing in RNA Structure and Function
The base pairing rules, both canonical and non-canonical, dictate the secondary and tertiary structures of RNA molecules. These structures are intimately linked to their biological functions.
mRNA (Messenger RNA)
mRNA carries the genetic information from DNA to the ribosomes, where protein synthesis takes place. While mRNA primarily exists as a linear molecule, its secondary structure, influenced by base pairing, can impact its stability, translation efficiency, and interactions with other molecules. Hairpin loops and other secondary structure elements can affect mRNA translation initiation and termination.
tRNA (Transfer RNA)
tRNA molecules are adapter molecules that carry amino acids to the ribosome during protein synthesis. Their highly specific three-dimensional structure, largely determined by base pairing (both canonical and wobble), is critical for their interaction with mRNA codons and aminoacyl-tRNA synthetases. The anticodon loop, formed through base pairing, recognizes specific codons on mRNA.
rRNA (Ribosomal RNA)
rRNA constitutes the structural and catalytic core of ribosomes. The intricate base pairing interactions within rRNA molecules create a complex three-dimensional structure essential for its catalytic activity in peptide bond formation during protein synthesis. The precise arrangement of bases and the resultant structure are key to ribosomal function.
snRNA (Small Nuclear RNA)
snRNAs (small nuclear RNAs) are involved in pre-mRNA splicing, a crucial step in eukaryotic gene expression. Their base-pairing interactions with pre-mRNA and other proteins are essential for accurate intron excision and exon ligation.
miRNA (Micro RNA)
miRNAs are short RNA molecules that regulate gene expression by binding to target mRNAs and inhibiting their translation or promoting their degradation. Base pairing between the miRNA and its target mRNA is crucial for this gene regulatory function.
Techniques for Studying RNA Base Pairing
Several techniques allow researchers to study RNA base pairing and structure:
- X-ray crystallography: Provides high-resolution structural information, revealing the precise arrangement of bases and other atoms within RNA molecules.
- NMR spectroscopy: Another powerful technique that provides information about RNA structure and dynamics in solution.
- Computational methods: Various computational approaches predict RNA secondary and tertiary structures based on the base pairing rules and energy minimization principles. These methods are invaluable for analyzing large RNA sequences and studying the effects of mutations.
Implications of Base Pairing in Disease
Disruptions in RNA base pairing, either through mutations or environmental factors, can have profound consequences, leading to various diseases. For example:
- Mutations affecting tRNA structure: Can lead to impaired protein synthesis and various genetic disorders.
- Disruptions in mRNA secondary structure: Can affect translation efficiency and protein production.
- Alterations in rRNA structure: Can compromise ribosomal function and overall protein synthesis.
Conclusion
The base pairing rule for RNA, while fundamentally similar to DNA, exhibits significant variations that significantly impact RNA's diverse functionalities. The canonical A-U and G-C base pairs are the cornerstone of RNA secondary structure formation, but the inclusion of wobble base pairing and various non-canonical interactions expands the structural and functional repertoire of RNA molecules. Understanding these rules and their exceptions is crucial for comprehending the intricate roles RNA plays in cellular processes and the molecular basis of various diseases. Further research continues to unravel the complexities of RNA structure and function, leading to significant advancements in various fields, including medicine and biotechnology. The continued exploration of RNA base pairing will undoubtedly reveal further insights into the fundamental processes of life and pave the way for novel therapeutic interventions.
Latest Posts
Latest Posts
-
Expected Number Of Trials Until Success
Apr 06, 2025
-
Determinants Of The Elasticity Of Supply
Apr 06, 2025
-
What Does A Replicated Chromosome Look Like
Apr 06, 2025
-
How To Find The Sum Of An Alternating Series
Apr 06, 2025
-
Magnetic Field From North To South
Apr 06, 2025
Related Post
Thank you for visiting our website which covers about What Is The Base Pairing Rule For Rna . We hope the information provided has been useful to you. Feel free to contact us if you have any questions or need further assistance. See you next time and don't miss to bookmark.
